The biopharmaceutical industry is undergoing a profound transformation as artificial intelligence (AI) shifts antibody discovery and development from trial-and-error to a predictable and programmable engineering process [1]. By leveraging advanced AI and machine learning models, researchers can now rapidly analyze massive biological datasets to simulate complex molecular interactions, predict developability attributes, and co-optimize multiple drug-like properties. Furthermore, generative AI is beginning to enable the de novo design of novel antibodies tailored to specific targets or epitopes [2, 3]. As AI moves from point solutions to end-to-end systems, the greatest impact emerges when learning, design, and experimental validation are tightly coupled.
To translate AI innovation into tangible outcomes, Nona Biosciences developed Hu-mAtrIx™, a next-generation AI-powered platform that seamlessly embeds closed-loop learning into antibody discovery. Hu-mAtrIx™ is proactively updated by Nona with each experimental cycle, enabling rapid, data-driven optimization of antibody sequence and function. As a result, the platform is poised as the next-generation engine for fully human therapeutic antibodies, delivering compressed discovery timelines, reduced development costs, improved developability, and higher success rates across therapeutic antibody programs.
AI-Integrated Antibody Discovery Workflow
A persistent challenge in antibody discovery is the high failure rate of initial candidates. Many antibodies initially identified in discovery campaigns might exhibit suboptimal biophysical properties, including low solubility, aggregation, chemical instability, immunogenic potential, or challenges in manufacturing scale-up [4, 5, 6]. When these liabilities are not identified during early discovery, they often surface later, contributing directly to CMC delays, costly re-engineering efforts, or program termination.
In this context, developability, encompassing the features that determine a binder’s fitness for drug development, including efficacy, pharmacology, safety, and manufacturability, has become a central concept in antibody discovery [7]. Modern antibody developability assessment integrates analytical screening with wet-lab validated predictive AI tools across the antibody pipeline, enabling more informed hit selection, earlier risk identification, and reduced downstream attrition.
Antibody discovery typically begins with the screening of thousands of antibody candidates recovered from immunization or display (e.g., phage display) campaigns. Following sequence elucidation, enabled by next-generation sequencing (NGS), computational/AI-based models are increasingly applied to predict functional binding-related features (e.g., affinity ranking, paratope/epitope relationships, or polyreactivity), and developability liabilities (e.g., aggregation hotspots, viscosity risk, chemical instability, or immunogenicity) directly from the antibody sequences [4, 6, 8].
These sequence-level predictions enable early triaging of large hit sets, substantially reducing the number of candidates advanced to recombinant expression and downstream biological evaluation (i.e., binding and functional assays). Importantly, predicted liabilities are not used solely as exclusion criteria. Instead, they inform rational antibody engineering strategies by highlighting sequence or structural motifs that may be targeted for engineering, such as posttranslational modification (PTMs) sites or unfavorable charged and hydrophobic patches within complementarity determining regions (CDRs) [6].
High-throughput AI-Integrated Antibody Discovery Workflow
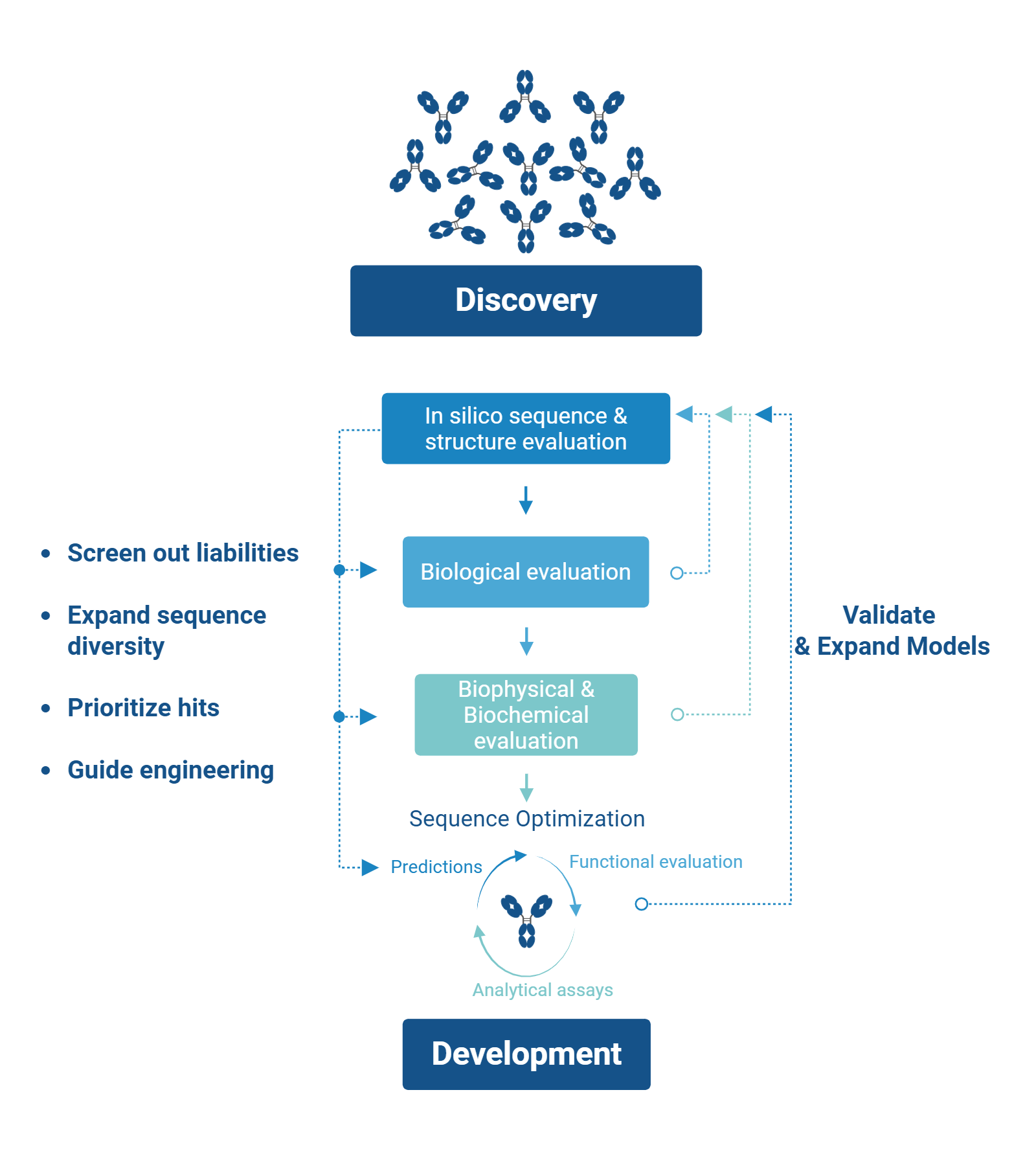
As candidates progress, biophysical and biochemical characterization focuses on identifying liabilities related to stability, colloidal behavior, specificity, expression yield, and immunogenicity risk. Together, insights gathered from computational prediction, functional assays, and analytical measurements guide downstream optimization strategies [6, 7].
Once engineering strategies are implemented, functional and analytical assessments must be repeated to experimentally confirm that the predicted liabilities have been mitigated without compromising binding or biological activity. Iterative experimental confirmation not only advances improved candidates but also strengthens the underlying predictive models through closed-loop feedback.
From Developability Concepts to Measurable Risk
Assessing developability in vitro
Conventional antibody developability assessment involves the systematic evaluation of biophysical, biochemical, and manufacturability attributes to estimate the likelihood that a binder can progress successfully from discovery through clinical development and commercial production. While developability is often discussed conceptually, in practice, it is evaluated through defined, measurable properties that correlate with downstream CMC, formulation, and clinical risk.
The field has converged on a core panel of stress conditions and standardized analytical methods designed to reveal liabilities that commonly emerge during scale‑up, storage, or manufacturing [6]. Many of these risks, such as aggregation, self‑interaction, chemical instability, or polyspecific binding, are not readily apparent from binding data alone, yet they account for a substantial proportion of late‑stage attrition.
The table below provides a high‑level view of the key developability attributes typically evaluated during early antibody discovery, the types of stress or degradation conditions used to reveal hidden risks, and the corresponding predictive analytical screening approaches that inform CMC and advancement decisions.
Analytical Methods for Antibody Developability Assessment
| Developability Attribute | Key Risk Assessed | Stress / Screening Context | Primary Assays | Impact on Development Decisions |
|---|---|---|---|---|
| Expression / Yield | Manufacturability and secretion efficiency | Transient/stable expression in HEK293 or CHO | Small-scale expression; Protein A yield; SEC | Cost of goods, platform compatibility, early attrition |
| Structural Stability | Resistance to unfolding and conformational change | Thermal and pH stress | DSF; DSC | Shelf life, formulation robustness |
| Chemical Stability / PTM | Susceptibility to oxidation, deamidation, and glycation | Oxidative, thermal, pH stress, and forced glycation | LC-MS peptide mapping; intact mass | Potency drift, degradation control |
| Aggregation Propensity | Formation of soluble/insoluble aggregates | Heat, agitation, freeze-thaw | SEC; DLS | Immunogenicity risk, yield loss |
| Colloidal Stability / Self-Interaction | Self-association and viscosity risk | High concentration; formulation stress | AC-SINS; DLS | SC delivery feasibility, high-dose formulation |
| Fragmentation Risk | Backbone cleavage (hinge, CH2–CH3) | Elevated temperature; pH extremes | CE-SDS; SEC; LC-MS | Potency retention, immunogenicity |
| Hydrophobicity Risk | Exposed hydrophobic surfaces | High-salt stress (e.g., ammonium sulfate) | HIC | Aggregation risk, nonspecific interactions |
| Charge Heterogeneity | Acidic/basic variants impacting PK | pH and ionic strength variation | cIEF; IEX | Purification fit, formulation strategy |
| Polyspecificity | Off-target binding and clearance risk | Native conditions | Polyreactivity ELISA; polyspecificity panels | PK predictability, safety margin |
| Solubility Risk | Maximum achievable concentration | High-concentration screening | Turbidity; visual solubility | Dose and route of administration |
Affinity-capture self-interaction nanoparticle spectroscopy (AC-SINS), capillary electrophoresis (CE-SDS), capillary isoelectric focusing (cIEF), dynamic light scattering (DLS), differential scanning calorimetry (DSC), differential scanning fluorimetry (DSF), hydrophobic interaction chromatography (HIC), ion-exchange chromatography (IEX), liquid chromatography mass spectrometry (LC-MS), size-exclusion chromatography (SEC). Adapted and simplified from Bailly et al., 2020 [6]. This overview highlights the intent behind each measurement, what risk it is designed to uncover, and why it matters for progression decisions.
Assessing developability in silico
Among AI applications in antibody discovery, developability prediction offers particularly high downstream impact because it directly targets the biophysical and manufacturability risks responsible for many late-stage failures. Models trained on large datasets linking sequence and structure to experimental outcomes can predict many liabilities before extensive wet‑lab screening begins [7].
Computational and AI-Based Antibody Developability Assessment from Sequence and Structure
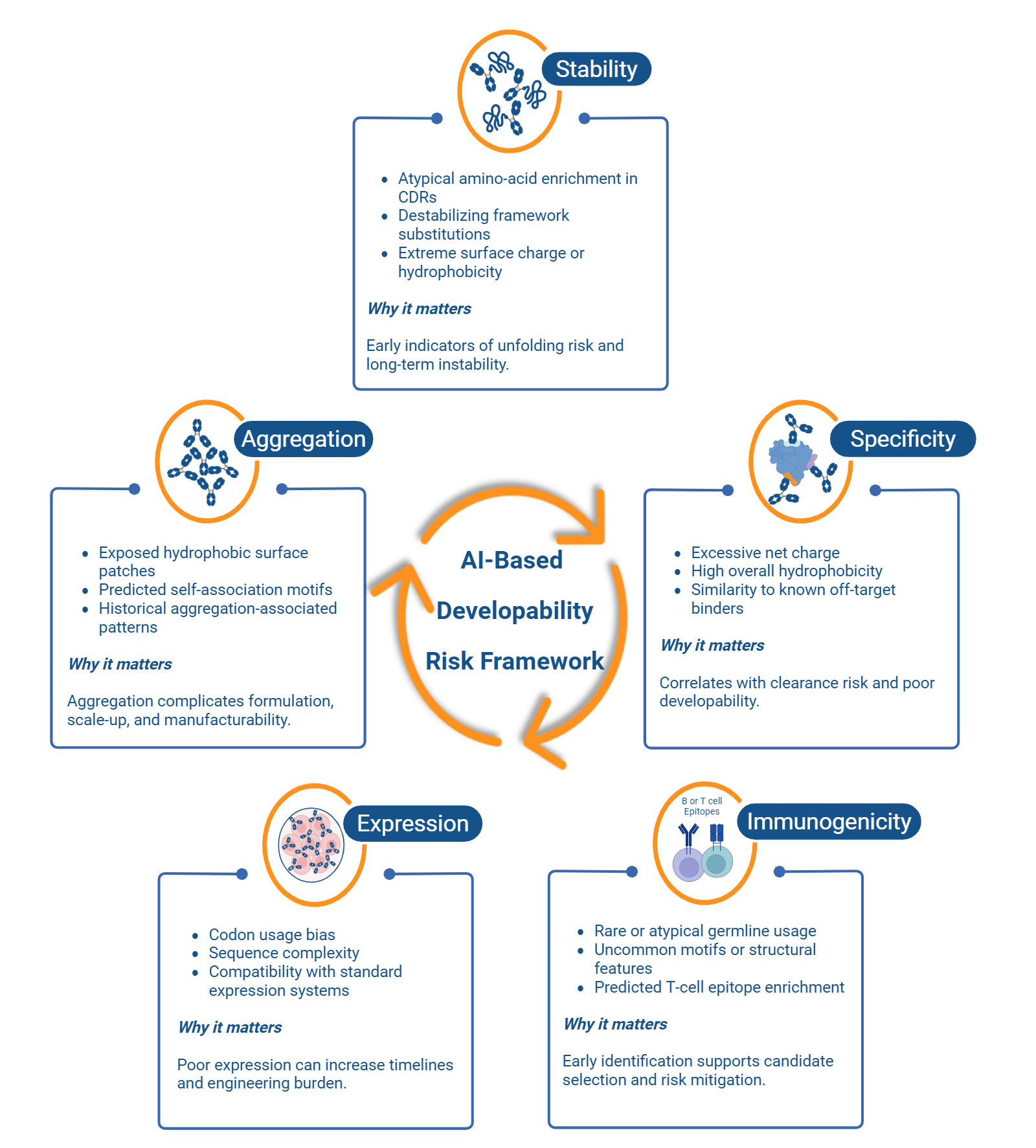
When deployed as decision-support tools rather than replacements for experimental validation, AI-based developability models allow discovery teams to focus resources on candidates that combine strong biological activity with a higher likelihood of downstream success. Their utility depends on careful benchmarking against high quality experimental data and continuous calibration through iterative feedback [6, 8].
AI-Integrated Antibody Discovery at Nona Biosciences
What is HumAtrIx™?
Hu-mAtrIx™ is Nona Bioscience’s proprietary AI platform designed to support antibody discovery and early optimization within Harbour Mice®-based programs (i.e., fully human H2L2 and HCAb generating Harbour Mice). Rather than replacing established and clinically proven discovery workflows, Hu-mAtrIx™ functions as an integrated analytical layer, applying data-driven insights at key decision points to support candidate selection, prioritization, and risk mitigation. By embedding AI directly into in vivo discovery programs, Hu-mAtrIx™ ensures that predictive insights remain anchored in clinically relevant, experimentally validated antibody repertoires. This integration preserves biological diversity while improving the efficiency and consistency of early-stage decision making.
Data Foundation
The predictive performance of Hu-mAtrIx™ is driven by a combination of proprietary and public datasets, including:
- Large-scale, fully human HCAb antibody libraries comprising millions of sequences
- Public protein and antibody sequence datasets spanning tens of millions of entries
- Target-specific binding and functional data generated directly from H2L2 and HCAb Harbour Mice® discovery campaigns
Together, these data enable Hu-mAtrIx™ to capture both broad sequence-level patterns and target-specific binding characteristics, supporting robust prediction across diverse discovery programs.
Integrated Capabilities
Hu-mAtrIx™ incorporates multiple AI-driven analytical modules applied across early discovery and optimization, rather than relying on a single predictive function.
Hu-mAtrIx™ Integration within the Harbour Mice® Antibody Discovery Workflow
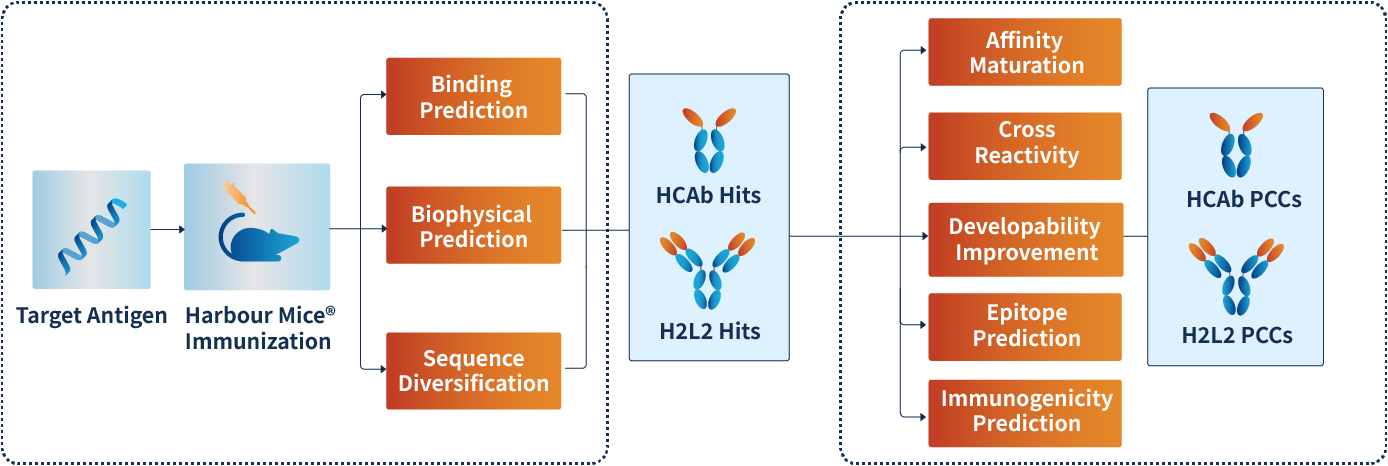
Guided lead prioritization
Machine-learning models support ranking and prioritization of antibody sequences based on predicted binding potential, diversity, and early biophysical indicators such as self-interaction, aggregation tendency, and thermal or colloidal stability.
Predictive affinity assessment
Sequence-based models provide early insight into relative binding performance, helping Nona’s team determine which candidates to advance, deprioritize, or engineer before committing to resource-intensive experimental optimization.
Developability and risk prediction
AI models within Hu-mAtrIx™ assess key developability attributes, including molecular stability, specificity, and immunogenicity risk. Early identification of these liabilities allows for targeted mitigation before candidates enter advanced optimization or IND-enabling workflows.
Sequence diversification
Hu-mAtrIx™ also incorporates generative protein language models to support sequence diversification alongside in vivo discovery. These models are designed to expand biologically relevant sequence space while remaining tightly coupled to experimentally validated repertoires and wet-lab feedback. Generated sequences serve as exploratory inputs rather than replacements for empirical discovery.
Placement within Nona’s discovery workflow
Hu-mAtrIx™ contributes AI-driven insights at multiple stages of antibody discovery:
- In parallel with NGS, expanding discovery space using data from prior Harbour Mice® campaigns
- During post-NGS candidate triage, enabling efficient prioritization of large sequence pools
- At hit-to-lead transition points, informing advancement decisions
- During early optimization, guiding affinity and developability tradeoffs before extensive wet-lab investment
At each stage, the platform is designed to reduce experimental burden while improving the quality and consistency of advancement decisions.

An Integrated Approach to AI in Discovery
Hu-mAtrIx™ reflects Nona Biosciences’ broader approach to applying AI in biologics discovery: deploying machine learning where it delivers clear value while grounding predictions in high-quality biological data. By functioning as an integrated analytical partner rather than a standalone solution, Hu-mAtrIx™ supports more informed decision-making, accelerates discovery timelines, and helps translate antibody diversity into development-ready candidates without compromising biological relevance.
Related Resources
To explore AI-integrated antibody discovery and developability optimization in greater detail:
- Download our Brochure on Hu-mAtrIx™ for an overview of how our AI-powered platform accelerates fully human antibody discovery through predictive modeling, closed-loop learning, and data-driven optimization.
- View our Scientific Poster on De Novo Fully Human VH Antibody Discovery for a concise, visual summary of AI-driven sequence generation, developability prediction, and experimental validation within Harbour Mice®–based workflows.